DNA Subway
A fun and easy-to-use interface for learning, teaching, and doing high-level genome analysis.
DNA Subway is an educational bioinformatics platform that bundles research-grade bioinformatics tools, high-performance computing, and databases into genomics workflows. Designed for teaching and exploration, it combines research-grade software, high-performance computing, and genomics databases.

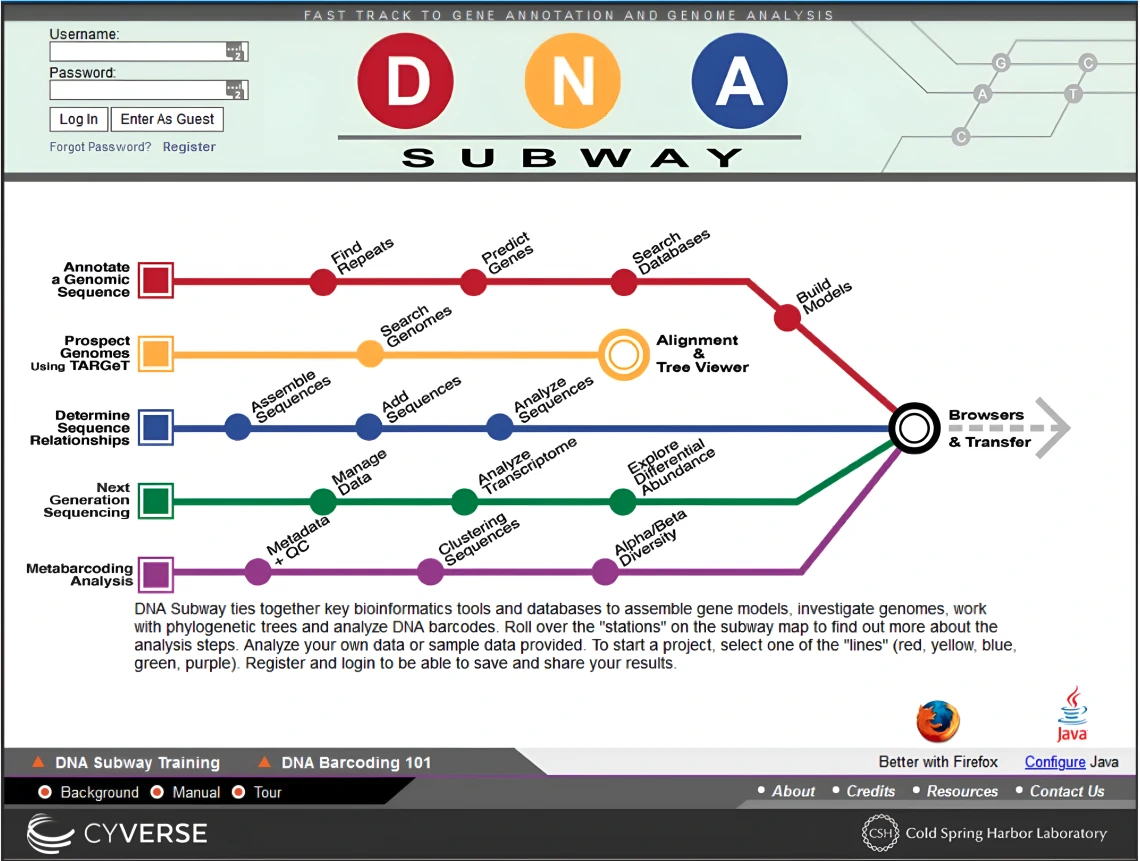
Users can choose from five color-coded “Subway Lines,” each representing a different type of analysis:
(Red Line) – Predict and annotate genes (up to 150 kb of DNA)
(Yellow Line) – Analyze homologs in sequenced genomes
(Blue Line) – Identify species using DNA barcoding and phylogenetic trees
(Green Line) – Examine RNA-Seq datasets for differential transcript abundance
(Purple Line) – Explore eDNA and metabarcoding data using QIIME
Hop on and take your data for a ride!
Getting Started
- Teach students (or learn yourself!) to analyze DNA using bioinformatics tools
- Predict and annotate genes in up to 150,000 base pairs of DNA sequence (Red Line)
- Prospect entire genomes for related genes and sequences (Yellow Line)
- Determine sequence relationships, view phylogenetic trees, and analyze "DNA barcodes" (Blue Line)
- Analyze RNA-Seq data for differential expression (GreenLine)
- Analyze microbiome and eDNA using QIIME2 (Purple Line)
Key Features
Modular bioinformatics workflows (subway lines) composed of research-grade, open-source tools for genomics, transcriptomics, and metagenomics analyses.
User-friendly graphical interface that lowers the barrier to entry for executing complex analyses such as gene prediction, sequence alignment, phylogenetic inference, and differential expression.
Support for custom and public data—users can import their own sequence files or utilize curated reference datasets integrated into the platform.
Companion educational resources available by (DNA Barcoding 101), including lab protocols, curriculum-aligned project guides, and scaffolding for DNA barcoding experiments.
Documentation

