When
Where
Webinar Materials
Jacob's Tutorial and Slides in GitHub
About the Webinar
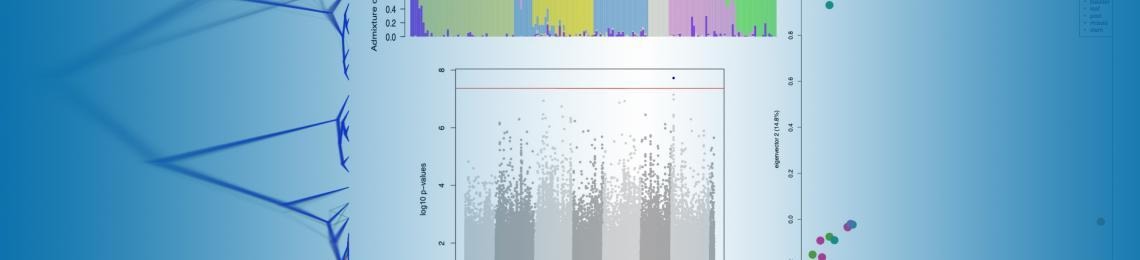
Following up on the earlier popular webinar he gave in Fall 2020 on SNP Calling, Jacob Landis, of the Boyce Thompson Institute's Computational Biology Center, will highlight a few of the downstream PopGen and Evolutionary analyses you can do after you have generated variant calls in your study system. If you have data that shows variants (differences between individuals or between populations), this webinar will provide useful steps using CyVerse to handle a variety of different data types, such as RNA-Seq, RAD-Seq, Hyb-Seq, or genome resequencing, to measure isolation by distance, population structure, genome wide association (GWAS) and inference on evolutionary relatedness.
What You'll Learn
- Steps to do downstream analyses on a variety of different variants data types
- How to use and where to find CyVerse resources to do downstream PopGen and Evolution studies on variants
Suggested Expertise Level
- Intermediate genomics and coding skills are helpful
- For background and basics, review Jacob's previous webinar on SNP Calling
About the Presenter

Jacob Landis is a Lecturer at Cornell University's School of Integrative Plant Science and working out of the Specht Lab. He received his doctorate in Botany from University of Florida, Gainsville, and now focuses his research in Evolutionary Genomics on the genomic changes associated with adaptation in barley. Like most scientists, he is involved with several projects, including studying rice diversity in Surinam and floral development across monocots.

